Create Phylogenetic Tree of SARS CoV 2 by UShER
There are nearly 14 million viral genome sequences right now in the GISAID EpiCoV ™ database. It is not likely to infer the phylogenetic relationships for such a huge dataset by traditional maximum likelyhood or Bayesian methods in a shor time period. The UShER package was developed to generate ultra-large phylogenetic tree of SARS-CoV-2 genomes. The algorithm of the UShER program is to place new samples onto an existing phylogeny using maximum parsimony method. It is able to place given SARS-CoV-2 genome sequences into the GISAID global phylogeny in a couple of hours. This program is particularly helpful in understanding the relationships of newly sequenced SARS-CoV-2 genomes with each other and with previously sequenced genomes in a global phylogeny.
The UShER package is composed by four programs:
UShER: a program that rapidly places new samples onto an existing phylogeny using maximum parsimony.matUtils: a toolkit for querying, interpreting and manipulating the mutation-annotated trees (MATs).matOptimize: a program to rapidly and effectively optimize a mutation-annotated tree (MAT) for parsimony using subtree pruning and regrafting (SPR) moves within a user-defined radius.RIPPLES: a program that uses a phylogenomic technique to rapidly and sensitively detect recombinant nodes and their ancestors in a mutation-annotated tree (MAT).
The taxoniumtools and Taxonium website are used to display the MAT generated by UShER.
1. Installation
1.1 Install UShER package via conda
# Create a new environment for UShER
conda create -n usher
# Activate the newly created environment
conda activate usher
# Set up channels
conda config --add channels defaults
conda config --add channels bioconda
conda config --add channels conda-forge
# Install the UShER package
conda install usher
1.2 Install taxoniumtools
Although the UShER wiki recommends matUtils extract to produce a MAT for Taxonium visualization, Taxonium recommends taxoniumtools to do this conversion for current Taxonium Version 2.
# If Python 3.x were not installed in the usher environment
# It will also install pip
conda install -c conda-forge python
# Install taxoniumtools
pip install taxoniumtools
2. Procedures
2.1 Download sequences and metadata
Download public tree (pb format), sample sequences, reference genome, and sites to mask:
# Protocol buffer format Mutation Annotated Tree, generated at 2021-08-04
wget http://hgdownload.soe.ucsc.edu/goldenPath/wuhCor1/UShER_SARS-CoV-2/2021/08/04/public-2021-08-04.all.masked.pb
# Metadata
wget https://hgdownload.soe.ucsc.edu/goldenPath/wuhCor1/UShER_SARS-CoV-2/2021/08/04/public-2021-08-04.metadata.tsv.gz
# 15 sample sequences, S1 ~ S15
wget https://raw.githubusercontent.com/bpt26/usher_wiki/main/docs/source/test_samples.fa
# Reference genome, NC_045512v2
wget https://hgdownload.soe.ucsc.edu/goldenPath/wuhCor1/bigZips/wuhCor1.fa.gz
# GenBank format format annotation file of reference genome
wget https://raw.githubusercontent.com/theosanderson/taxonium/master/taxoniumtools/test_data/hu1.gb
# Problem sites, will be masked
wget https://raw.githubusercontent.com/W-L/ProblematicSites_SARS-CoV2/master/problematic_sites_sarsCov2.vcf
Note:
- The latest protcol buffer format Mutation Annotated Tree (MAT) is available at:
http://hgdownload.soe.ucsc.edu/goldenPath/wuhCor1/UShER_SARS-CoV-2/public-latest.all.masked.pb.gz- This tree includes only public sequences aggregated from GenBank, COG-UK, and the China National Center for Bioinformation. Please read more details at:
2.2 Operation
- Unzip reference sequence:
gunzip wuhCor1.fa.gz
- Alignment sample sequences against reference genome by MAFFT
mafft --thread 10 --auto --keeplength --addfragments test_samples.fa wuhCor1.fa > aligned_seqs.fa
Note:
MAFFT options:
--auto: automatically switches algorithm according to data size. Safer to always use this flag.--keeplength: the alignment length is unchanged. Insertions at the fragmentary sequences are deleted.--addfragments: Adding unaligned fragmentary sequence(s) into an existing alignment.
- Convert above FASTA format alignment file to Variant Call Format (VCF) single-nucleotide diffs
faToVcf -maskSites=problematic_sites_sarsCov2.vcf aligned_seqs.fa aligned_seqs.vcf
- Now, add test samples to the protobuf MAT:
usher -T 10 -i public-latest.all.masked.pb.gz -v aligned_seqs.vcf -o user_seqs.pb
- Convert UShER
pbformat MAT to Taxoniumjsnlformat:
usher_to_taxonium --input user_seqs.pb --output user_seqs-taxonium.jsonl.gz --metadata public-2021-08-04.metadata.tsv.gz --genbank hu1.gb --columns genbank_accession,country,date,pangolin_lineage
Note: Do not download gene annotation file from NC_045512.2 GenBank record directly, use the optimized annotation file
hu1.gbfrom taxonium.
2.3 Tree visualization
Open the generated user_seqs-taxonium.jsonl.gz tree file at Taxonium website.
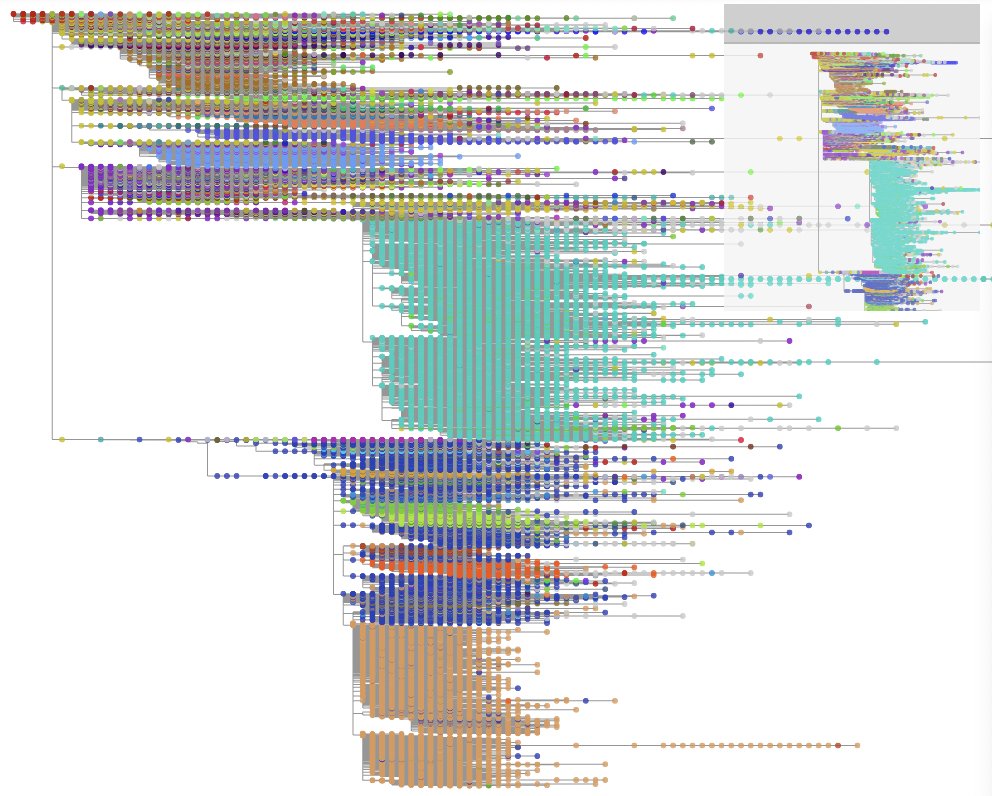
Please see the Taxonium documentation for usage details.
References
Appendix A. Command line options
A.1 UShER help
UShER (v0.6.0)
Options:
-v [ --vcf ] arg Input VCF file (in uncompressed or
gzip-compressed .gz format) [REQUIRED]
-t [ --tree ] arg Input tree file
-d [ --outdir ] arg (=.) Output directory to dump output and log
files [DEFAULT uses current directory]
-i [ --load-mutation-annotated-tree ] arg
Load mutation-annotated tree object
-o [ --save-mutation-annotated-tree ] arg
Save output mutation-annotated tree
object to the specified filename
-s [ --sort-before-placement-1 ] Sort new samples based on computed
parsimony score and then number of
optimal placements before the actual
placement [EXPERIMENTAL].
-S [ --sort-before-placement-2 ] Sort new samples based on the number of
optimal placements and then the
parsimony score before the actual
placement [EXPERIMENTAL].
-A [ --sort-before-placement-3 ] Sort new samples based on the number of
ambiguous bases [EXPERIMENTAL].
-r [ --reverse-sort ] Reverse the sorting order of sorting
options (sort-before-placement-1 or
sort-before-placement-2) [EXPERIMENTAL]
-c [ --collapse-tree ] Collapse internal nodes of the input
tree with no mutations and condense
identical sequences in polytomies into
a single node and the save the tree to
file condensed-tree.nh in outdir
-C [ --collapse-output-tree ] Collapse internal nodes of the output
tree with no mutations before the
saving the tree to file final-tree.nh
in outdir
-e [ --max-uncertainty-per-sample ] arg (=1000000)
Maximum number of equally parsimonious
placements allowed per sample beyond
which the sample is ignored
-E [ --max-parsimony-per-sample ] arg (=1000000)
Maximum parsimony score of the most
parsimonious placement(s) allowed per
sample beyond which the sample is
ignored
-u [ --write-uncondensed-final-tree ]
Write the final tree in uncondensed
format and save to file
uncondensed-final-tree.nh in outdir
-k [ --write-subtrees-size ] arg (=0) Write minimum set of subtrees covering
the newly added samples of size equal
to this value
-K [ --write-single-subtree ] arg (=0)
Similar to write-subtrees-size but
produces a single subtree with all
newly added samples along with random
samples up to the value specified by
this argument
-p [ --write-parsimony-scores-per-node ]
Write the parsimony scores for adding
new samples at each existing node in
the tree without modifying the tree in
a file names parsimony-scores.tsv in
outdir
-M [ --multiple-placements ] arg (=1) Create a new tree up to this limit for
each possibility of parsimony-optimal
placement
-l [ --retain-input-branch-lengths ] Retain the branch lengths from the
input tree in out newick files instead
of using number of mutations for the
branch lengths.
-n [ --no-add ] Do not add new samples to the tree
-D [ --detailed-clades ] In clades.txt, write a histogram of
annotated clades and counts across all
equally parsimonious placements
-T [ --threads ] arg (=104) Number of threads to use when possible
[DEFAULT uses all available cores, 104
detected on this machine]
--version Print version number
-h [ --help ] Print help messages
A.2 usher_to_taxonium help
usage: usher_to_taxonium [-h] -i INPUT -o OUTPUT [-m METADATA] [-g GENBANK]
[-c COLUMNS] [-C]
[--chronumental_steps CHRONUMENTAL_STEPS]
[--chronumental_date_output CHRONUMENTAL_DATE_OUTPUT]
[--chronumental_tree_output CHRONUMENTAL_TREE_OUTPUT]
[--chronumental_reference_node CHRONUMENTAL_REFERENCE_NODE]
[-j CONFIG_JSON] [-t TITLE]
[--overlay_html OVERLAY_HTML] [--remove_after_pipe]
[--clade_types CLADE_TYPES] [--name_internal_nodes]
[--shear] [--shear_threshold SHEAR_THRESHOLD]
[--only_variable_sites] [--key_column KEY_COLUMN]
Convert a Usher pb to Taxonium jsonl format
options:
-h, --help show this help message and exit
-i INPUT, --input INPUT
File path to input Usher protobuf file (.pb)
-o OUTPUT, --output OUTPUT
File path for output Taxonium jsonl file
-m METADATA, --metadata METADATA
File path for input metadata file (CSV/TSV)
-g GENBANK, --genbank GENBANK
File path for GenBank file containing reference genome
(N.B. currently only one chromosome is supported, and
no compound features)
-c COLUMNS, --columns COLUMNS
Column names to include in the metadata, separated by
commas, e.g. `pangolin_lineage,country`
-C, --chronumental Runs Chronumental to build a time tree. The metadata
TSV must include a date column.
--chronumental_steps CHRONUMENTAL_STEPS
Number of steps to run Chronumental for
--chronumental_date_output CHRONUMENTAL_DATE_OUTPUT
Optional output file for the chronumental date table
if you want to keep it (a table mapping nodes to their
inferred dates).
--chronumental_tree_output CHRONUMENTAL_TREE_OUTPUT
Optional output file for the chronumental time tree
file in nwk format.
--chronumental_reference_node CHRONUMENTAL_REFERENCE_NODE
A reference node to be used for Chronumental. This
should be earlier in the outbreak and have a good
defined date. If not set the oldest sample will be
automatically picked by Chronumental.
-j CONFIG_JSON, --config_json CONFIG_JSON
A JSON file to use as a config file containing things
such as search parameters
-t TITLE, --title TITLE
A title for the tree. This will be shown at the top of
the window as "[Title] - powered by Taxonium"
--overlay_html OVERLAY_HTML
A file containing HTML to put in the About box when
this tree is loaded. This could contain information
about who built the tree and what data you used.
--remove_after_pipe If set, we will remove anything after a pipe (|) in
each node's name, after joining to metadata
--clade_types CLADE_TYPES
Optionally specify clade types provided in the UShER
file, comma separated - e.g. 'nextstrain,pango'. Order
must match that used in the UShER pb file. If you
haven't specifically annotated clades in your
protobuf, don't use this
--name_internal_nodes
If set, we will name internal nodes node_xxx
--shear If set, we will 'shear' the tree. This will iterate
over all nodes. If a particular sub-branch makes up
fewer than e.g. 1/1000 of the total descendants, then
in most cases it represents a sequencing error. (But
it also could represent recombinants, or a real, unfit
branch.) We remove these to simplify the
interpretation of the tree.
--shear_threshold SHEAR_THRESHOLD
Threshold for shearing, default is 1000 meaning
branches will be removed if they make up less than
<1/1000 nodes. Has no effect unless --shear is set.
--only_variable_sites
Only store information about the root sequence at a
particular position if there is variation at that
position somewhere in the tree. This helps to speed up
the loading of larger genomes such as MPXV.
--key_column KEY_COLUMN
The column in the metadata file which is the same as
the names in the tree
A.3 faToVcf help
faToVcf - Convert a FASTA alignment file to Variant Call Format (VCF) single-nucleotide diffs
usage:
faToVcf in.fa out.vcf
options:
-ambiguousToN Treat all IUPAC ambiguous bases (N, R, V etc) as N (no call).
-excludeFile=file Exclude sequences named in file which has one sequence name per line
-includeNoAltN Include base positions with no alternate alleles observed, but at
least one N (missing base / no-call)
-includeRef Include the reference in the genotype columns
(default: omitted as redundant)
-maskSites=file Exclude variants in positions recommended for masking in file
(typically https://github.com/W-L/ProblematicSites_SARS-CoV2/raw/master/problematic_sites_sarsCov2.vcf)
-maxDiff=N Exclude sequences with more than N mismatches with the reference
(if -windowSize is used, sequences are masked accordingly first)
-minAc=N Ignore alternate alleles observed fewer than N times
-minAf=F Ignore alternate alleles observed in less than F of non-N bases
-minAmbigInWindow=N When -windowSize is provided, mask any base for which there are at
least this many N, ambiguous or gap characters within the window.
(default: 2)
-noGenotypes Output 8-column VCF, without the sample genotype columns
-ref=seqName Use seqName as the reference sequence; must be present in faFile
(default: first sequence in faFile)
-resolveAmbiguous For IUPAC ambiguous characters like R (A or G), if the character
represents two bases and one is the reference base, convert it to the
non-reference base. Otherwise convert it to N.
-startOffset=N Add N bases to each position (for trimmed alignments)
-vcfChrom=seqName Use seqName for the CHROM column in VCF (default: ref sequence)
-windowSize=N Mask any base for which there are at least -minAmbigWindow bases in a
window of +-N bases around the base. Masking approach adapted from
https://github.com/roblanf/sarscov2phylo/ file scripts/mask_seq.py
Use -windowSize=7 for same results.
in.fa must contain a series of sequences with different names and the same length.
Both N and - are treated as missing information.